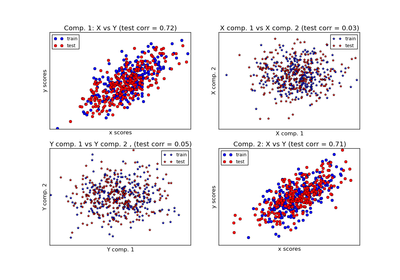
sklearn.cross_decomposition.PLSCanonical¶
- class sklearn.cross_decomposition.PLSCanonical(n_components=2, scale=True, algorithm='nipals', max_iter=500, tol=1e-06, copy=True)[source]¶
PLSCanonical implements the 2 blocks canonical PLS of the original Wold algorithm [Tenenhaus 1998] p.204, referred as PLS-C2A in [Wegelin 2000].
This class inherits from PLS with mode=”A” and deflation_mode=”canonical”, norm_y_weights=True and algorithm=”nipals”, but svd should provide similar results up to numerical errors.
Read more in the User Guide.
Parameters: scale : boolean, scale data? (default True)
algorithm : string, “nipals” or “svd”
The algorithm used to estimate the weights. It will be called n_components times, i.e. once for each iteration of the outer loop.
max_iter : an integer, (default 500)
the maximum number of iterations of the NIPALS inner loop (used only if algorithm=”nipals”)
tol : non-negative real, default 1e-06
the tolerance used in the iterative algorithm
copy : boolean, default True
Whether the deflation should be done on a copy. Let the default value to True unless you don’t care about side effect
n_components : int, number of components to keep. (default 2).
Attributes: x_weights_ : array, shape = [p, n_components]
X block weights vectors.
y_weights_ : array, shape = [q, n_components]
Y block weights vectors.
x_loadings_ : array, shape = [p, n_components]
X block loadings vectors.
y_loadings_ : array, shape = [q, n_components]
Y block loadings vectors.
x_scores_ : array, shape = [n_samples, n_components]
X scores.
y_scores_ : array, shape = [n_samples, n_components]
Y scores.
x_rotations_ : array, shape = [p, n_components]
X block to latents rotations.
y_rotations_ : array, shape = [q, n_components]
Y block to latents rotations.
n_iter_ : array-like
Number of iterations of the NIPALS inner loop for each component. Not useful if the algorithm provided is “svd”.
Notes
Matrices:
T: x_scores_ U: y_scores_ W: x_weights_ C: y_weights_ P: x_loadings_ Q: y_loadings__
Are computed such that:
X = T P.T + Err and Y = U Q.T + Err T[:, k] = Xk W[:, k] for k in range(n_components) U[:, k] = Yk C[:, k] for k in range(n_components) x_rotations_ = W (P.T W)^(-1) y_rotations_ = C (Q.T C)^(-1)
where Xk and Yk are residual matrices at iteration k.
Slides explaining PLS <http://www.eigenvector.com/Docs/Wise_pls_properties.pdf>
For each component k, find weights u, v that optimize:
max corr(Xk u, Yk v) * std(Xk u) std(Yk u), such that ``|u| = |v| = 1``
Note that it maximizes both the correlations between the scores and the intra-block variances.
The residual matrix of X (Xk+1) block is obtained by the deflation on the current X score: x_score.
The residual matrix of Y (Yk+1) block is obtained by deflation on the current Y score. This performs a canonical symmetric version of the PLS regression. But slightly different than the CCA. This is mostly used for modeling.
This implementation provides the same results that the “plspm” package provided in the R language (R-project), using the function plsca(X, Y). Results are equal or collinear with the function pls(..., mode = "canonical") of the “mixOmics” package. The difference relies in the fact that mixOmics implementation does not exactly implement the Wold algorithm since it does not normalize y_weights to one.
References
Jacob A. Wegelin. A survey of Partial Least Squares (PLS) methods, with emphasis on the two-block case. Technical Report 371, Department of Statistics, University of Washington, Seattle, 2000.
Tenenhaus, M. (1998). La regression PLS: theorie et pratique. Paris: Editions Technic.
Examples
>>> from sklearn.cross_decomposition import PLSCanonical >>> X = [[0., 0., 1.], [1.,0.,0.], [2.,2.,2.], [2.,5.,4.]] >>> Y = [[0.1, -0.2], [0.9, 1.1], [6.2, 5.9], [11.9, 12.3]] >>> plsca = PLSCanonical(n_components=2) >>> plsca.fit(X, Y) ... PLSCanonical(algorithm='nipals', copy=True, max_iter=500, n_components=2, scale=True, tol=1e-06) >>> X_c, Y_c = plsca.transform(X, Y)
Methods
fit(X, Y) Fit model to data. fit_transform(X[, y]) Learn and apply the dimension reduction on the train data. get_params([deep]) Get parameters for this estimator. predict(X[, copy]) Apply the dimension reduction learned on the train data. score(X, y[, sample_weight]) Returns the coefficient of determination R^2 of the prediction. set_params(**params) Set the parameters of this estimator. transform(X[, Y, copy]) Apply the dimension reduction learned on the train data. - __init__(n_components=2, scale=True, algorithm='nipals', max_iter=500, tol=1e-06, copy=True)[source]¶
- fit(X, Y)[source]¶
Fit model to data.
Parameters: X : array-like, shape = [n_samples, n_features]
Training vectors, where n_samples in the number of samples and n_features is the number of predictors.
Y : array-like of response, shape = [n_samples, n_targets]
Target vectors, where n_samples in the number of samples and n_targets is the number of response variables.
- fit_transform(X, y=None, **fit_params)[source]¶
Learn and apply the dimension reduction on the train data.
Parameters: X : array-like of predictors, shape = [n_samples, p]
Training vectors, where n_samples in the number of samples and p is the number of predictors.
Y : array-like of response, shape = [n_samples, q], optional
Training vectors, where n_samples in the number of samples and q is the number of response variables.
copy : boolean, default True
Whether to copy X and Y, or perform in-place normalization.
Returns: x_scores if Y is not given, (x_scores, y_scores) otherwise. :
- get_params(deep=True)[source]¶
Get parameters for this estimator.
Parameters: deep: boolean, optional :
If True, will return the parameters for this estimator and contained subobjects that are estimators.
Returns: params : mapping of string to any
Parameter names mapped to their values.
- predict(X, copy=True)[source]¶
Apply the dimension reduction learned on the train data.
Parameters: X : array-like of predictors, shape = [n_samples, p]
Training vectors, where n_samples in the number of samples and p is the number of predictors.
copy : boolean, default True
Whether to copy X and Y, or perform in-place normalization.
Notes
This call requires the estimation of a p x q matrix, which may be an issue in high dimensional space.
- score(X, y, sample_weight=None)[source]¶
Returns the coefficient of determination R^2 of the prediction.
The coefficient R^2 is defined as (1 - u/v), where u is the regression sum of squares ((y_true - y_pred) ** 2).sum() and v is the residual sum of squares ((y_true - y_true.mean()) ** 2).sum(). Best possible score is 1.0 and it can be negative (because the model can be arbitrarily worse). A constant model that always predicts the expected value of y, disregarding the input features, would get a R^2 score of 0.0.
Parameters: X : array-like, shape = (n_samples, n_features)
Test samples.
y : array-like, shape = (n_samples) or (n_samples, n_outputs)
True values for X.
sample_weight : array-like, shape = [n_samples], optional
Sample weights.
Returns: score : float
R^2 of self.predict(X) wrt. y.
- set_params(**params)[source]¶
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as pipelines). The former have parameters of the form <component>__<parameter> so that it’s possible to update each component of a nested object.
Returns: self :
- transform(X, Y=None, copy=True)[source]¶
Apply the dimension reduction learned on the train data.
Parameters: X : array-like of predictors, shape = [n_samples, p]
Training vectors, where n_samples in the number of samples and p is the number of predictors.
Y : array-like of response, shape = [n_samples, q], optional
Training vectors, where n_samples in the number of samples and q is the number of response variables.
copy : boolean, default True
Whether to copy X and Y, or perform in-place normalization.
Returns: x_scores if Y is not given, (x_scores, y_scores) otherwise. :